
You have generated three transgenic lines of maize that are resistant to the European corn borer, a significant pest in many regions of the world. The transgenic lines (T
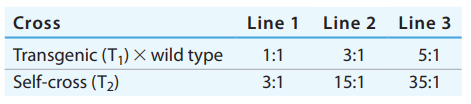
Explain these segregation ratios.
Want to see the full answer?
Check out a sample textbook solution
Chapter 15 Solutions
Genetic Analysis: An Integrated Approach (3rd Edition)
Additional Science Textbook Solutions
Essentials of Genetics (9th Edition) - Standalone book
Campbell Essential Biology with Physiology (6th Edition)
Laboratory Experiments in Microbiology (12th Edition) (What's New in Microbiology)
Microbiology with Diseases by Taxonomy (5th Edition)
Biology: Life on Earth (11th Edition)
- Your TA gives you an Escherichia coli strain (AmreB) that carries a gene deletion in the mreB gene. As a result, the strain is not able to produce the MreB protein. Your task is to compare the morphology of actively growing AmreB cells to the parental wildtype strain (produces MreB). What difference in morphology will you likely observe? Would you expect the same in Staphylococcus aureus?arrow_forwardYour supervisor asked you to insert a specific stress-tolerant gene into a tissue cultured plant batch. What can be your possible approach? You may only give a flow chart to show your process.arrow_forwardT. Miyake and M. Demerec examined proline-requiring mutations in the bacterium Salmonella typhimurium (). On the basis of complementation testing, they found four proline auxotrophs: proA, proB, proC, and proD. To determine whether proA, proB, proC, and proD loci were located close together on the bacterial chromosome, they conducted a transduction experiment. Bacterial strains that were proC+ and had mutations at proA, proB, or proD were used as donors. The donors were infected with bacteriophages, and progeny phages were allowed to infect recipient bacteria with genotype proC− proA+ proB+ proD+. The recipient bacteria werethen plated on a selective medium that allowed only proC+ bacteria to grow. After this, the proC+ transductants were plated on selective media to reveal their genotypes at the other three pro loci. The following results were obtained: Q.Why are there no proC− genotypes among the transductants?arrow_forward
- T. Miyake and M. Demerec examined proline-requiring mutations in the bacterium Salmonella typhimurium (). On the basis of complementation testing, they found four proline auxotrophs: proA, proB, proC, and proD. To determine whether proA, proB, proC, and proD loci were located close together on the bacterial chromosome, they conducted a transduction experiment. Bacterial strains that were proC+ and had mutations at proA, proB, or proD were used as donors. The donors were infected with bacteriophages, and progeny phages were allowed to infect recipient bacteria with genotype proC− proA+ proB+ proD+. The recipient bacteria werethen plated on a selective medium that allowed only proC+ bacteria to grow. After this, the proC+ transductants were plated on selective media to reveal their genotypes at the other three pro loci. The following results were obtained: Q.Is there evidence that proA, proB, and proD are located close to proC? Explain your answer.arrow_forwardT. Miyake and M. Demerec examined proline-requiring mutations in the bacterium Salmonella typhimurium (). On the basis of complementation testing, they found four proline auxotrophs: proA, proB, proC, and proD. To determine whether proA, proB, proC, and proD loci were located close together on the bacterial chromosome, they conducted a transduction experiment. Bacterial strains that were proC+ and had mutations at proA, proB, or proD were used as donors. The donors were infected with bacteriophages, and progeny phages were allowed to infect recipient bacteria with genotype proC− proA+ proB+ proD+. The recipient bacteria werethen plated on a selective medium that allowed only proC+ bacteria to grow. After this, the proC+ transductants were plated on selective media to reveal their genotypes at the other three pro loci. The following results were obtained: Q.Which genotypes represent single transductants and which represent cotransductants?arrow_forwardTransposon mutagenesis was used to generate a library of mutants within the Salmonella genome. You are trying to identify a colony with the transposon inserted in the pathogenic related gene SPI-1 using PCR. Forward and reverse primers are generated that flank either side of the gene and yield a wild type product that is 900 bases in length. Which of the colonies sampled in the gel would you expect to contain the SPI-1 gene with transposon insertion? 3,000 2,000 1,000 700 500 300 100 Ladder Colony A Colony B Colony C Colony D Colony E none colonies A&C colonies B&E O colonies A, C, &D colonies B, D, &E -arrow_forward
- You are cloning a gene called ice, which will help strawberry plants survive in cold weather, into a test group of strawberry plants. Select the events you would use to genetically engineer these plants. Insert the ice gene into the T-DNA region of a Ti plasmid; transform Agrobacterium tumefaciens with this plasmid; Inoculate a culture of strawberry plant cells with Agrobacterium tumefaciens; Select plant cells that have taken up the ice gene. Insert the ice gene into the T-DNA region of the Agrobacterium tumefaciens chromosome; Inoculate a culture of strawberry plant cells with Agrobacterium tumefaciens; Select plant cells that have taken up the ice gene. O Insert the ice gene into a Ti transposon; Introduce the transposon into Agrobacterium tumefaciens Ti plasmid: Inoculate a culture of strawberry plant cells with Agrobacterium tumefaciens: Select plant cells that have taken up the ice gene. Insert the ice gene into a Ti bacteriophage; transduce Agrobacterium tumefaciens with this…arrow_forwardDNA from a strain of bacteria with genotype a+ b+ c+ d+ e+ was isolated and used to transform a strain of bacteria that was a− b− c− d− e−. The transformants were tested for the presence of donated genes. The following genes were cotransformed: a+ and d+ b+ and e+ c+ and d+ c+ and e+ What is the order of genes a, b, c, d, and e on the bacterial chromosome?arrow_forwardpZERO®-1 is a 2808 bp cloning vector from Invitrogen. This vector allows effective selection of positive recombinants via disruption of the lethal gene, ccdB. Besides the lethal gene, it has an additional Zeocin resistance gene for tighter screening control. The Zeocin resistance gene in this vector is certainly more advantageous over ampicillin resistance gene. Give its THREE (3) advantages.arrow_forward
- Cystic fibrosis (CF) is a genetic disorder affecting a number of organs, including the lung airways, pancreas, and sweat glands. Mutations in both copies of the CFTR gene causes cystic fibrosis. Imagine that you have sweat gland samples from several Cystic Fibrosis patients (A-C) with unknown mutations in CFTR. You also have normal (+) sweat gland sample to use as a positive control. А В С А В С Choose which mutation would explain the RNA and protein results in A, B, & C: 1. Promoter/Regulatory mutation 2. Silent mutation 3. Missense mutation 4. Deletion mutation 5. Splice site mutation 6. Nonsense mutation RNA gel Protein gelarrow_forwardA research group is studying a bacterium X that binds to mucosal cells in the lung and invades. Wildtype X has an LD50 value of 10 bacteria when administered to mice by inhalation. Using transposon mutagenesis, the researchers have isolated two mutants of X that they call Xmut1 and Xmut2, both of which have LD50 values of 105 when inhaled by mice. However, in tissue culture cells, Xmut1 can invade the cells just as well as wild-type X, while Xmut2 cannot. Provide a possible explanation for these results.arrow_forwardBacteriophage P22 was used in generalised transduction experiments to infect the Salmonella typhimurium donor strains described in the table below. The resulting phage lysates were then used to infect the recipient strains of S. typhimurium recipient strains listed in the table. In each cross, a phenotype was selected for one of the selected for one of the three genetic markers studied (str, aceA, thrA), and were made to select the recombinants corresponding to the other two markers. markers. The results are given in the following table: Strain I donor str thrA aceA thrA str aceA+ Strain recipient strs thrA+ aceA thrA str aceA Phenotype selected Str Ace+ Str recombinants selected ThrA ThrA ThrA ThrA Ace Ace Number 60 40 95 5 10 90 str: gene involved in streptomycin resistance, aceA: gene involved in the use of acetate as a carbon source, thrA: gene involved in threonine biosynthesis. 1) What are the selective media used in these three transduction experiments? to obtain the selected…arrow_forward
 Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON
Human Anatomy & Physiology (11th Edition)BiologyISBN:9780134580999Author:Elaine N. Marieb, Katja N. HoehnPublisher:PEARSON Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax
Biology 2eBiologyISBN:9781947172517Author:Matthew Douglas, Jung Choi, Mary Ann ClarkPublisher:OpenStax Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education,
Anatomy & PhysiologyBiologyISBN:9781259398629Author:McKinley, Michael P., O'loughlin, Valerie Dean, Bidle, Theresa StouterPublisher:Mcgraw Hill Education, Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company
Molecular Biology of the Cell (Sixth Edition)BiologyISBN:9780815344322Author:Bruce Alberts, Alexander D. Johnson, Julian Lewis, David Morgan, Martin Raff, Keith Roberts, Peter WalterPublisher:W. W. Norton & Company Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co.
Laboratory Manual For Human Anatomy & PhysiologyBiologyISBN:9781260159363Author:Martin, Terry R., Prentice-craver, CynthiaPublisher:McGraw-Hill Publishing Co. Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education
Inquiry Into Life (16th Edition)BiologyISBN:9781260231700Author:Sylvia S. Mader, Michael WindelspechtPublisher:McGraw Hill Education





